Simulation Engines
Cell Illustrator (CI) allows researchers to simulate complex biological processes. The modeling and simulation engine of Cell Illustrator is based on an extension of the Petri-net methodology. Cell Illustrator offers two modes of simulation described below.
Standard Simulation Engine
The standard simulation engine is dedicated for creating and testing of simulation models. It is fully integrated within the CI workspace window and allows for interactive pathway simulation. The CI user can:
- specify mathematical formulas for biochemical reactions
- define complex logic of the Petri-net by writing scripts
- test/run the simulation inside the CI workspace in an interactive way
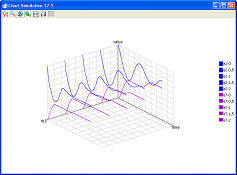
- view/compare the simulation results on charts
- save the simulation log and then visualize it in Cell Illustrator Player
SECG - Simulation Engine Code Generator
SECG is based on the same HFPNe model as the standard simulation engine described above, however it is faster, more customizable and adds special viewing and running modes. The SECG idea is to generate source code for the simulated pathway model. This source code can be compiled and executed within the Cell Illustrator workspace or in an external program or framework.
SECG can be used to:
- simulate large models.
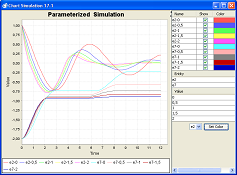
- run parameterized simulations - perform a serious of simulations of the same model with varying values of selected parameter(s)
- compare the results of parameterized simulations in order to obtain the optimum values for the parameters
- export the simulation model to a programming language, e.g. Java, C, C++, Fortran, Perl, Python
 |
 |
Scripting Languages. Compatibility
Scripts are used in Cell Illustrator to define complex logic of the Petri Net. The scripting languages are different in the versions 4.0 and 5.0 of Cell Illustrator. Cell Illustrator 5.0 supports several scripting languages: simplemath, Java, JavaScript (js) and Pnuts, while Cell Illustrator 4.0 supports one language only - Pnuts. In rare cases it can happen that models created with Cell Illustrator 4.0 cannot be simulated in the version 5.0. The document Writing and Porting Scripts in Cell Illustrator 5.0 describes how to port such CSML models created with CI 4.0 and run simulations in CI 5.0.

